Differences between version 265 and predecessor to the previous major change of HomePage.
Other diffs: Previous Revision, Previous Author
| Newer page: |
version 265 |
Last edited on 04 December 2018 17:43 |
by Emilio Martinez Nuñez |
Revert |
| Older page: |
version 223 |
Last edited on 05 July 2018 5:14 |
by Emilio Martinez Nuñez |
Revert |
@@ -1,48 +1,43 @@
-{{colorbox
-|text
=<big>
**Transition State Search Using Chemical Dynamics Simulations**</big>
-|color
=silver
-|bordercolor
=white
-}}
+==
**Transition State Search Using Chemical Dynamics Simulations** ==
-This program package has been designed to discover reaction mechanisms and solve the kinetics in an automated fashion, using chemical dynamics simulations. The basic idea is to obtain transition state (TS) guess structures from trajectory simulations performed at very high energies or temperatures. From the obtained TS structures, minima and product fragments are determined following the intrinsic reaction coordinate (IRC). Then, with all the stationary points, the reaction network is constructed. Finally, the kinetics is solved using the Kinetic Monte Carlo (KMC) method.
-The program is interfaced with MOPAC2016 and Gaussian 09 (G09), but work is in progress to incorporate more electronic structure programs.\\
+%color=red% (11/29/2018) A BUG THAT MIGHT CAUSE AN INFINITE LOOP IN THE KINETICS WAS FOUND
.%%
-Users are encouraged to read the tutorial (tutorial_tsscds.pdf) | http://forge.cesga.es/frs/?group_id
=77&release_id=21], which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA). The present version of the program can also be used to study homogeneous catalysis, but additional refinements are needed to make the code more general and user-friendly. This capability will be fully incorporated and described in the next release.\\
+%color
=red%PLEASE DOWNLOAD THE 2018rev1 VERSION WHERE THIS HAS BEEN CORRECTED%%
+[Download code (tsscds-SOURCE-2018rev1.tar.gz) & tutorial (tutorial_tsscds.pdf) | http://forge.cesga.es/frs/?group_id=77&release_id=21]\\
-The program has been tested so far on the following Linux distros: CentOS 7, Red Hat Enterprise Linux and Ubuntu 16.04.3 LTS
-====================AUTHORS==============================
-Emilio Martínez-Núñez\\
-Departamento de Química Física, Facultade de Química\\
-Avda. das Ciencias s/n\\
-15782 Santiago de Compostela, SPAIN\\
-mailto: emilio.nunez@usc.es
]
+[[Installation instructions]
]
-George L. Barnes\\
-Sabine Kopec\\
-Daniel Peláez\\
-Aurelio Rodríguez\\
-Roberto Rodríguez-Fernández\\
-James J. P. Stewart\\
-Saulo A. Vázquez
+[[How to cite the program]]
-====================DOWNLOADS==============================
-Download code
(tsscds-SOURCE-2018
.tar.gz
) & tutorial
(tutorial_tsscds
.pdf
) | http://forge
.cesga.es/frs/?group_id=77&release_id=21]
\\
+[[Research]]
+
+This program package has been designed to discover reaction mechanisms and solve the kinetics in an automated fashion, using chemical dynamics simulations. The basic idea is to obtain transition state
(TS) guess structures from trajectory simulations performed at very high energies or temperatures
. From the obtained TS structures, minima and product fragments are determined following the intrinsic reaction coordinate (IRC
). Then, with all the stationary points, the reaction network is constructed. Finally, the kinetics is solved using the Kinetic Monte Carlo
(KMC) method
.
+The program is interfaced with MOPAC2016 and Gaussian 09 (G09
), but work is in progress to incorporate more electronic structure programs
.\\
+
+[[Image:pr2.png]]
-Method
described in these two publications: [1 | http://onlinelibrary.wiley.com/doi/10.1002/jcc.23790/abstract]
+Details of the method are
described in these two publications: [1 | http://onlinelibrary.wiley.com/doi/10.1002/jcc.23790/abstract]
[2 | http://pubs.rsc.org/en/content/articlelanding/2015/cp/c5cp02175h#!divAbstract]
-=============================================================
-[[License]
]
+Users are encouraged to read the tutorial [ (tutorial_tsscds.pdf) | http://forge.cesga.es/frs/?group_id
=77&release_id
=21
], which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA).\\
-[[Installation instructions]]
-[[How to cite the
program]]
+The
program has been tested so far on the following Linux distros: CentOS 7, Red Hat Enterprise Linux and Ubuntu 16.04.3 LTS
-[[Research that employs tsscds]
]
+
+== Authors: ==
+
+George L. Barnes, Sabine Kopec, Daniel Peláez, Aurelio Rodríguez, Roberto Rodríguez-Fernández, James J. P. Stewart, Saulo A. Vázquez and //Emilio Martínez-Núñez//\\
+
+Departamento de Química Física\\
+Facultade de Química\\
+Avda. das Ciencias s/n\\
+15782 Santiago de Compostela, SPAIN\\
+
[mailto: emilio.nunez@usc.es
]
version 265
Transition State Search Using Chemical Dynamics Simulations
(11/29/2018) A BUG THAT MIGHT CAUSE AN INFINITE LOOP IN THE KINETICS WAS FOUND.
PLEASE DOWNLOAD THE 2018rev1 VERSION WHERE THIS HAS BEEN CORRECTED
 Download code (tsscds-SOURCE-2018rev1.tar.gz) & tutorial (tutorial_tsscds.pdf)
Download code (tsscds-SOURCE-2018rev1.tar.gz) & tutorial (tutorial_tsscds.pdf)
License
Installation instructions
Program execution
How to cite the program
Research
This program package has been designed to discover reaction mechanisms and solve the kinetics in an automated fashion, using chemical dynamics simulations. The basic idea is to obtain transition state (TS) guess structures from trajectory simulations performed at very high energies or temperatures. From the obtained TS structures, minima and product fragments are determined following the intrinsic reaction coordinate (IRC). Then, with all the stationary points, the reaction network is constructed. Finally, the kinetics is solved using the Kinetic Monte Carlo (KMC) method.
The program is interfaced with MOPAC2016 and Gaussian 09 (G09), but work is in progress to incorporate more electronic structure programs.
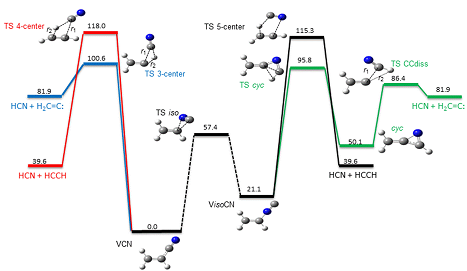
Details of the method are described in these two publications:  1
1
 2
2
Users are encouraged to read the tutorial  (tutorial_tsscds.pdf), which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA).
(tutorial_tsscds.pdf), which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA).
The program has been tested so far on the following Linux distros: CentOS 7, Red Hat Enterprise Linux and Ubuntu 16.04.3 LTS
Authors:
George L. Barnes, Sabine Kopec, Daniel Peláez, Aurelio Rodríguez, Roberto Rodríguez-Fernández, James J. P. Stewart, Saulo A. Vázquez and Emilio Martínez-Núñez
Departamento de Química Física
Facultade de Química
Avda. das Ciencias s/n
15782 Santiago de Compostela, SPAIN
 emilio.nunez@usc.es
emilio.nunez@usc.es
 Download code (tsscds-SOURCE-2018rev1.tar.gz) & tutorial (tutorial_tsscds.pdf)
Download code (tsscds-SOURCE-2018rev1.tar.gz) & tutorial (tutorial_tsscds.pdf)
 1
1
 2
2 (tutorial_tsscds.pdf), which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA).
(tutorial_tsscds.pdf), which is thought to guide you through the various steps necessary to predict reaction mechanisms and kinetics of unimolecular decompositions. To facilitate the presentation, we consider, as an example, the decomposition of formic acid (FA). emilio.nunez@usc.es
emilio.nunez@usc.es
